Integrating with MLflow¶
In this post, we are going to explain how setup the build-in integration
of sklearn-genetic-opt with MLflow.
To use this feature, we must set the parameters that will include
the tracking server, experiment, run name, tags and others,
the full implementation is here: MLflowConfig
Configuration¶
The configuration is pretty straight forward, we just need to import the main class and define some parameters, here there is its meaning:
tracking_uri: Address of local or remote tracking server.
experiment: Case sensitive name of an experiment to be activated.
run_name: Name of new run (stored as a mlflow.runName tag).
save_models: If
True, it will log the estimator into mlflow artifactsregistry_uri: Address of local or remote model registry server.
tags: Dictionary of tags to apply.
Example¶
In this example, we are going to log the information into a mlflow server that is running in our local host, port 5000, we want to save each of the trained models.
from sklearn_genetic.mlflow import MLflowConfig
mlflow_config = MLflowConfig(
tracking_uri="http://localhost:5000",
experiment="Digits-sklearn-genetic-opt",
run_name="Decision Tree",
save_models=True,
tags={"team": "sklearn-genetic-opt", "version": "0.5.0"})
Now, this config is passed to the GASearchCV class
in the parameter named log_config, for example:
from sklearn_genetic import GASearchCV
from sklearn_genetic.space import Categorical, Integer, Continuous
from sklearn.model_selection import train_test_split, StratifiedKFold
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import load_digits
from sklearn.metrics import accuracy_score
from sklearn_genetic.mlflow import MLflowConfig
data = load_digits()
label_names = data["target_names"]
y = data["target"]
X = data["data"]
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.33, random_state=42)
clf = DecisionTreeClassifier()
params_grid = {
"min_weight_fraction_leaf": Continuous(0, 0.5),
"criterion": Categorical(["gini", "entropy"]),
"max_depth": Integer(2, 20),
"max_leaf_nodes": Integer(2, 30)}
cv = StratifiedKFold(n_splits=3, shuffle=True)
evolved_estimator = GASearchCV(
clf,
cv=cv,
scoring="accuracy",
population_size=3,
generations=5,
tournament_size=3,
elitism=True,
crossover_probability=0.9,
mutation_probability=0.05,
param_grid=params_grid,
algorithm="eaMuPlusLambda",
n_jobs=-1,
verbose=True,
log_config=mlflow_config)
evolved_estimator.fit(X_train, y_train)
y_predict_ga = evolved_estimator.predict(X_test)
accuracy = accuracy_score(y_test, y_predict_ga)
print(evolved_estimator.best_params_)
Notice that we choose a small generations and population_size, just to be able to see the results without much verbosity.
If you go to you mlflow UI and click the experiment named “Digits-sklearn-genetic-opt” We should see something like this (I’ve hidden some columns to give a better look):
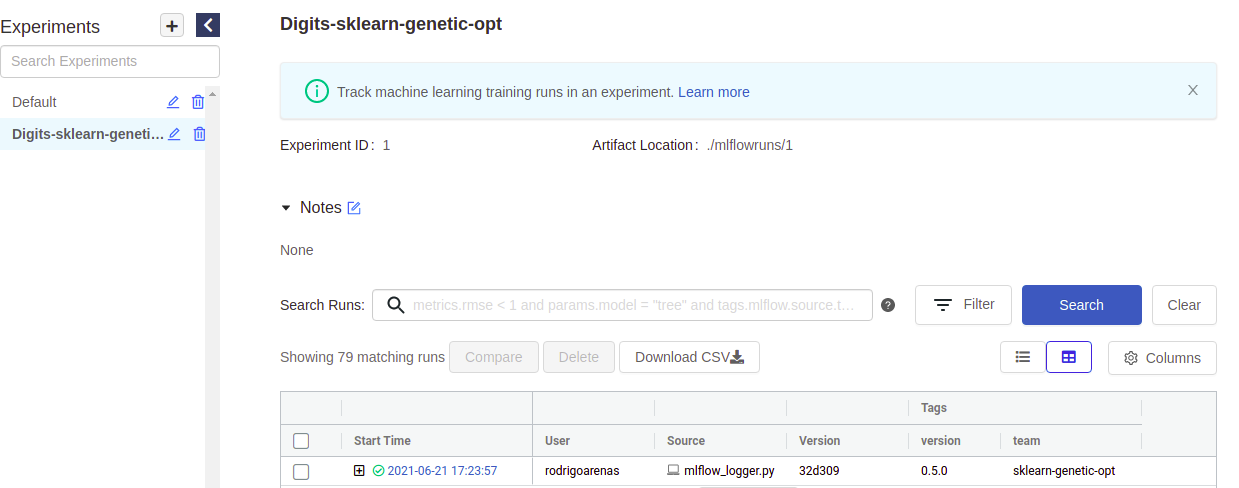
There we can see the user that ran the experiment, the name of the file which contained the source code, our tags and other metadata. Notice that there is “plus” symbol that will show us each of our iterations, this is because sklearn-genetic-opt will log each GASearchCV.fit() call in a nested way, think it like a parent run, and each child is one of the hyperparameters that were tested, for example if we run the same code again, now we see two parents runs:
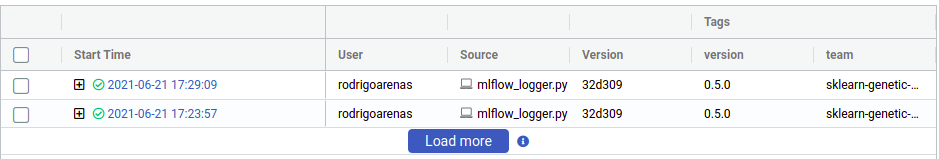
Now click on any of the “plus” symbols to see all the children, now they look like this (again edited the columns to display):
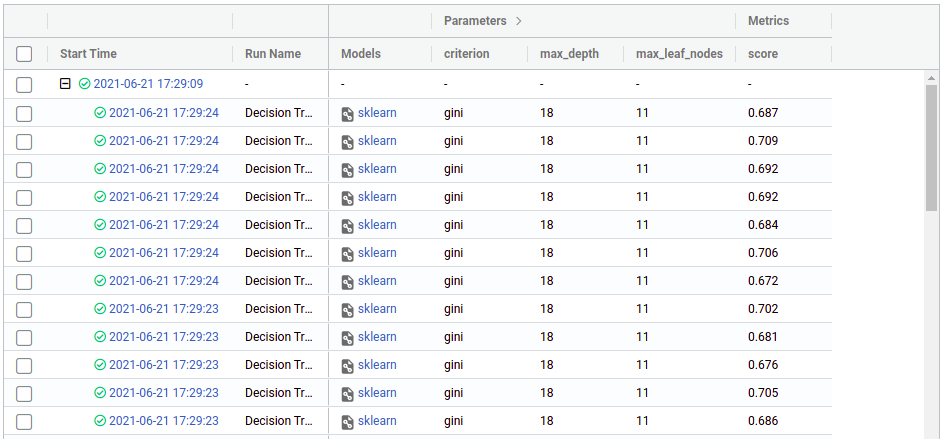
From there we can see the hyper parameters and the score (cross-validation) that we got in each run, from there we can use the regular mlflow functionalities like comparing runs, download the CSV, register a model, etc. You can see more on https://mlflow.org/docs/latest/index.html
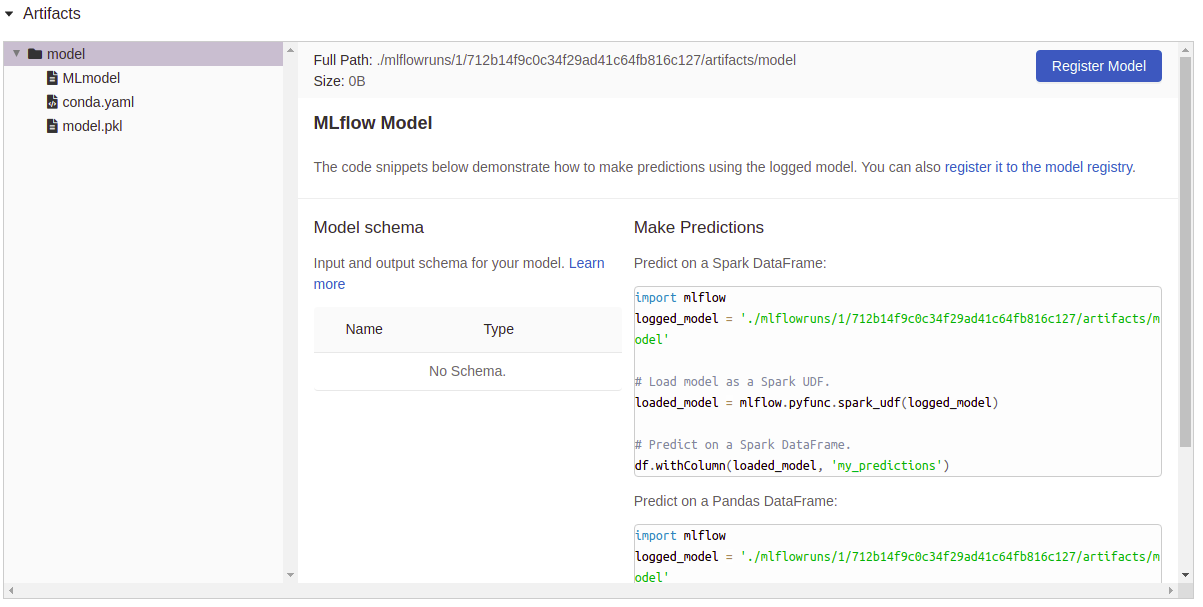
Now, as we set save_model=True, you can see that the column “Model”
as a file attached as an artifact, if we click on one of those, we see
a resume of that particular execution and some utils to use right away the
model: