Pipeline Prediction
[ ]:
import matplotlib.pyplot as plt
from sklearn_genetic import GASearchCV
from sklearn_genetic.space import Integer, Categorical, Continuous
from sklearn_genetic.plots import plot_fitness_evolution, plot_search_space
from sklearn_genetic.callbacks import LogbookSaver, ProgressBar
from sklearn.datasets import load_diabetes
from sklearn.model_selection import train_test_split, KFold
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.metrics import mean_squared_error
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
Import the data and split it in train and test sets
[2]:
data = load_diabetes()
y = data["target"]
X = data["data"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
Define the regressor to tune
[3]:
rg = GradientBoostingRegressor()
pipe = Pipeline([("scaler", StandardScaler()), ("rg", rg)])
Create the CV strategy and define the param grid
[4]:
cv = KFold(n_splits=5, shuffle=True)
param_grid = {
"rg__n_estimators": Integer(50, 600),
"rg__loss": Categorical(["absolute_error", "squared_error"]),
"rg__max_depth": Integer(2, 10),
"rg__learning_rate": Continuous(0.001, 0.01, distribution="log-uniform")}
Define the GASearchCV options
[5]:
evolved_estimator = GASearchCV(
estimator=pipe,
cv=cv,
scoring="neg_mean_squared_error",
population_size=8,
generations=15,
tournament_size=3,
elitism=True,
keep_top_k=4,
crossover_probability=0.9,
mutation_probability=0.08,
param_grid=param_grid,
criteria="max",
algorithm="eaMuCommaLambda",
n_jobs=-1)
Optionally, create some Callbacks
[6]:
callbacks = [LogbookSaver(checkpoint_path="./logbook.pkl")]
Fit the model and see some results
[7]:
evolved_estimator.fit(X_train, y_train, callbacks=callbacks)
y_predict_ga = evolved_estimator.predict(X_test)
mse = mean_squared_error(y_test, y_predict_ga)
gen nevals fitness fitness_std fitness_max fitness_min
0 8 -4238.26 565.058 -3756.78 -5523.09
1 16 -4051.7 352.254 -3756.29 -4631.79
2 16 -3665.2 70.1723 -3572.13 -3749.68
3 16 -3587.2 49.7041 -3507.56 -3644.64
4 16 -3513.45 102.322 -3420.14 -3698.88
5 16 -3678.71 164.066 -3521.83 -4011.67
6 15 -3547.3 35.2384 -3500.55 -3604.69
7 16 -3468.1 51.9507 -3379.15 -3542.45
8 16 -3472.2 51.3204 -3410.31 -3581.8
9 16 -3434.18 43.9115 -3371.14 -3514.54
10 16 -3410.84 94.4055 -3325.11 -3560.57
11 16 -3493.9 93.8796 -3393.02 -3662.27
12 16 -3506.98 32.4934 -3478.73 -3588.94
13 16 -3515.91 133.628 -3300.73 -3696.27
14 16 -3481.82 58.9704 -3399.49 -3616.37
15 16 -3468.11 10.7602 -3450.02 -3476.26
[8]:
print(evolved_estimator.best_params_)
print("mse: ", "{:.2f}".format(mse))
print("Best k solutions: ", evolved_estimator.hof)
{'rg__n_estimators': 561, 'rg__loss': 'squared_error', 'rg__max_depth': 3, 'rg__learning_rate': 0.006219524925263899}
mse: 3169.70
Best k solutions: {0: {'rg__n_estimators': 561, 'rg__loss': 'squared_error', 'rg__max_depth': 3, 'rg__learning_rate': 0.006219524925263899}, 1: {'rg__n_estimators': 504, 'rg__loss': 'squared_error', 'rg__max_depth': 3, 'rg__learning_rate': 0.006219524925263899}, 2: {'rg__n_estimators': 517, 'rg__loss': 'squared_error', 'rg__max_depth': 3, 'rg__learning_rate': 0.006219524925263899}, 3: {'rg__n_estimators': 561, 'rg__loss': 'squared_error', 'rg__max_depth': 3, 'rg__learning_rate': 0.004507098298712037}}
[9]:
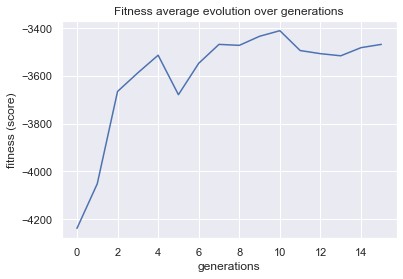
plot = plot_fitness_evolution(evolved_estimator, metric="fitness")
plt.show()
[10]:
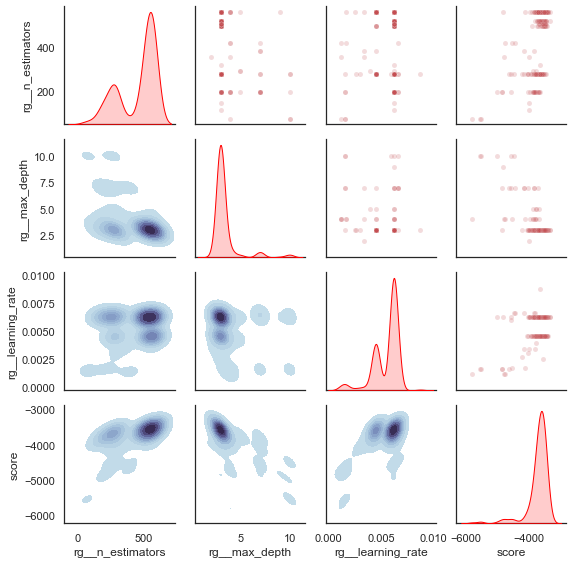
plot_search_space(evolved_estimator)
plt.show()
[ ]: