Iris Feature Selection
[ ]:
import matplotlib.pyplot as plt
from sklearn_genetic import GAFeatureSelectionCV
from sklearn_genetic.plots import plot_fitness_evolution
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
import numpy as np
Import the data and split it in train and test sets
Random noise is added to simulate useless variables
[2]:
data = load_iris()
X, y = data["data"], data["target"]
noise = np.random.uniform(0, 10, size=(X.shape[0], 10))
X = np.hstack((X, noise))
X.shape
[2]:
(150, 14)
Split the training and test data
[3]:
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=0)
Define the GAFeatureSelectionCV options
[4]:
clf = SVC(gamma='auto')
evolved_estimator = GAFeatureSelectionCV(
estimator=clf,
cv=3,
scoring="accuracy",
population_size=30,
generations=20,
n_jobs=-1,
verbose=True,
keep_top_k=2,
elitism=True,
)
Fit the model and see some results
[5]:
evolved_estimator.fit(X, y)
features = evolved_estimator.support_
# Predict only with the subset of selected features
y_predict_ga = evolved_estimator.predict(X_test)
accuracy = accuracy_score(y_test, y_predict_ga)
gen nevals fitness fitness_std fitness_max fitness_min
0 30 0.550444 0.153446 0.86 0.293333
1 60 0.636889 0.119365 0.86 0.473333
2 60 0.698667 0.11242 0.873333 0.46
3 60 0.707556 0.103876 0.873333 0.486667
4 60 0.723556 0.144086 0.9 0.366667
5 60 0.745556 0.152637 0.913333 0.366667
6 60 0.792889 0.108402 0.873333 0.513333
7 60 0.749111 0.16456 0.873333 0.413333
8 60 0.728889 0.179747 0.966667 0.373333
9 60 0.728222 0.158994 0.893333 0.42
10 60 0.785556 0.134892 0.94 0.48
11 60 0.733556 0.175942 0.94 0.44
12 60 0.784889 0.150554 0.94 0.413333
13 60 0.818444 0.148101 0.966667 0.413333
14 60 0.871778 0.116272 0.966667 0.453333
15 60 0.801556 0.184163 0.966667 0.386667
16 60 0.810222 0.163994 0.966667 0.393333
17 60 0.814222 0.148949 0.966667 0.44
18 60 0.72 0.182525 0.966667 0.366667
19 60 0.783556 0.156269 0.966667 0.42
20 60 0.803778 0.146694 0.966667 0.486667
[6]:
#Best features found
print(evolved_estimator.support_)
print("accuracy score: ", "{:.2f}".format(accuracy))
[False False True True False False False False False False False False
False False]
accuracy score: 0.96
[7]:
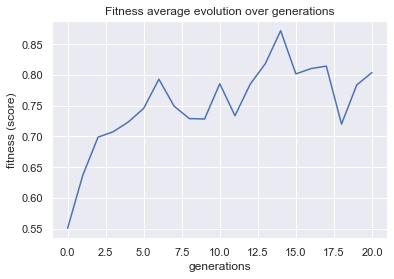
plot = plot_fitness_evolution(evolved_estimator, metric="fitness")
plt.show()
[8]:
# Convert the original input to the selected input
evolved_estimator.transform(X_test)
[8]:
array([[5.1, 2.4],
[4. , 1. ],
[1.4, 0.2],
[6.3, 1.8],
[1.5, 0.2],
[6. , 2.5],
[1.3, 0.3],
[4.7, 1.5],
[4.8, 1.4],
[4. , 1.3],
[5.6, 1.4],
[4.5, 1.5],
[4.7, 1.2],
[4.6, 1.5],
[4.7, 1.4],
[1.4, 0.1],
[4.5, 1.5],
[4.4, 1.2],
[1.4, 0.3],
[1.3, 0.4],
[4.9, 2. ],
[4.5, 1.5],
[1.9, 0.2],
[1.4, 0.2],
[4.8, 1.8],
[1. , 0.2],
[1.9, 0.4],
[4.3, 1.3],
[3.3, 1. ],
[1.6, 0.4],
[5.5, 1.8],
[4.5, 1.5],
[1.5, 0.2],
[4.9, 1.8],
[5.6, 2.2],
[3.9, 1.4],
[1.7, 0.3],
[5.1, 1.6],
[4.2, 1.5],
[4. , 1.2],
[5.5, 2.1],
[1.3, 0.2],
[5.1, 2.3],
[1.6, 0.6],
[1.5, 0.2],
[3.5, 1. ],
[5.5, 1.8],
[5.7, 2.5],
[5. , 1.5],
[5.8, 1.8]])
[ ]: